Case Study: GLP-1 and Incretin Mimetics with an AI Drug Discovery Partner
Kathy Wei¹; Baljit K. Ubhi²; Jack Howland²; Niall O’Connor²; Jennifer M. Campbell²; Geoffrey K. Feld³; Timothy P. Riley¹
¹AI drug discovery startup focusing on GLP-1 mimetics, San Francisco, CA | ²Matterworks, Inc., Somerville, MA | ³Geocyte LLC, Dublin, OH
Application: Screening AI-Designed Incretin Receptor Agonists
Peptide therapeutics targeting pancreatic incretin pathways have transformed the treatment of obesity and type 2 diabetes. However, limitations in metabolic stability, half-life, and multi-receptor selectivity constrain the potential of first-generation molecules. Scientists at an AI drug discovery startup focusing on GLP-1 mimetics engineered a suite of novel incretin mimetics designed to address these deficiencies, using a generative AI protein model to optimize structural and pharmacological properties.
A key candidate was a half-life extended (HLE) GLP-1 mimetic engineered to agonize multiple incretin receptors simultaneously, with the goal of improving both potency and dosing frequency. Its in vitro metabolic activity was evaluated alongside established control peptides (GLP-1, GIP, and dual GLP-1/GIP agonist GCG) using Pyxis-powered LC-MS metabolomics in a pancreatic cell model.
Experimental Details
Cells derived from a pancreatic adenocarcinoma cell line (CFPAC-1) were treated with each incretin mimetic at concentrations of 0.1 μM and 1.0 μM and incubated overnight. Following incubation, cell extracts and conditioned supernatants were prepared and analyzed by LC-MS using the standardized Pyxis method. Analytes were separated on a Waters Atlantis Premier BEH Z-HILIC column with a 6.5-minute gradient, and MS1 spectra were acquired in positive-negative switching mode on a ThermoFisher Scientific Orbitrap Exploris™ 120 mass spectrometer. Raw data files were uploaded to the Pyxis cloud platform for automated metabolite identification and quantitation.
Pyxis returned confident identities and concentrations for 135 biochemicals in cell extracts and 65 biochemicals in conditioned supernatants within 15 minutes of data upload, providing broad coverage of glycolysis, the TCA cycle, nucleotide metabolism, amino acid pathways, and the urea cycle.
Results: One-Day Functional Analysis Reveals Multiple Signatures of Drug Action
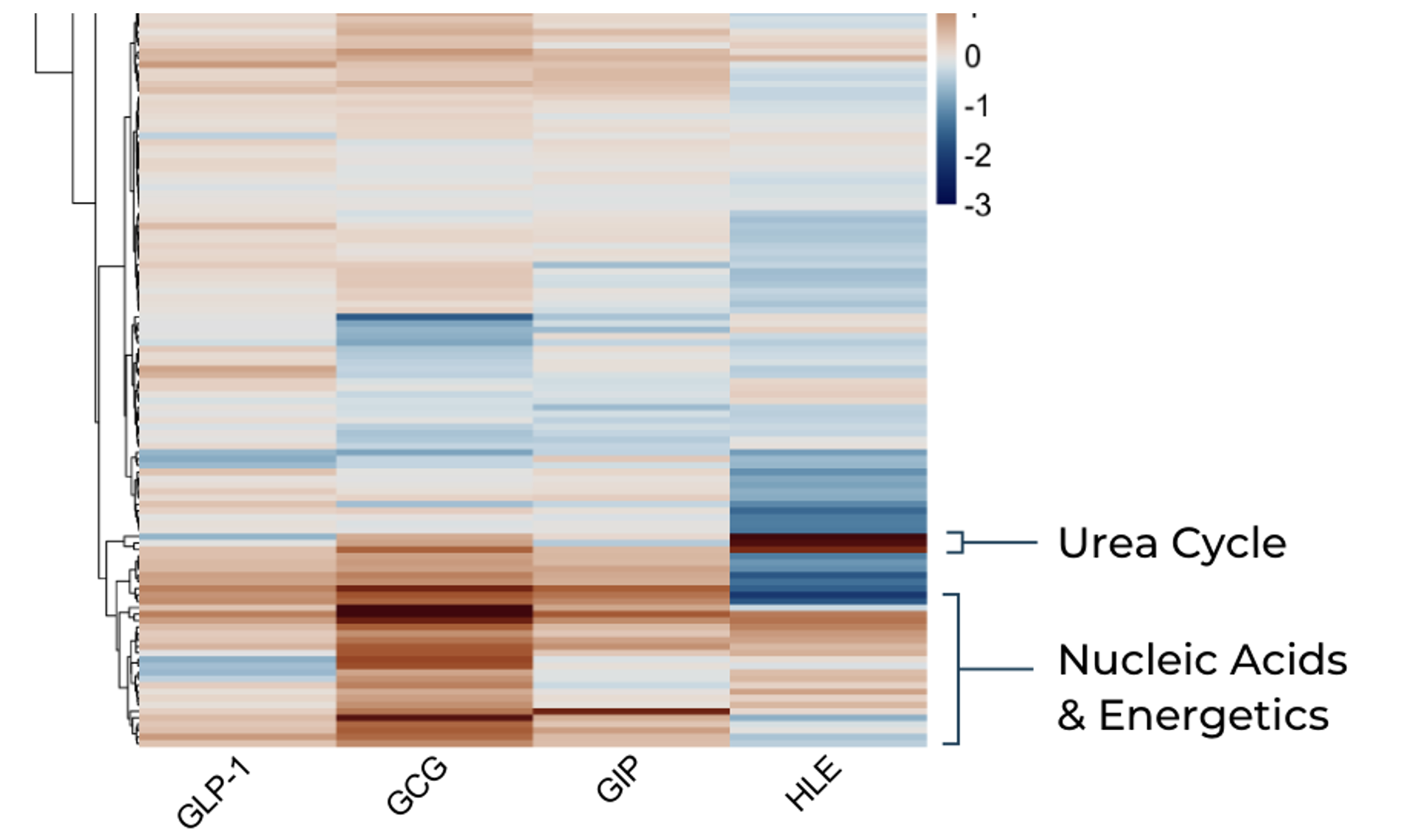
Hierarchical clustering analysis of cell extract metabolite profiles immediately revealed that the HLE novel mimetic induced a markedly distinct metabolic signature compared to the three control peptide mimetics (GLP-1, GIP, and GCG). While the control incretin receptor agonists clustered together, the HLE candidate separated clearly — a pattern that emerged without any a priori hypothesis about which pathways to interrogate.
Hierarchical clustering of cell extract metabolite profiles (1 μM drug vs. vehicle). The HLE GLP-1 mimetic (right column) induces a distinct metabolic pattern compared to GLP-1, GCG, and GIP controls, with prominent differences in urea cycle and nucleic acid/energetics pathway metabolites.
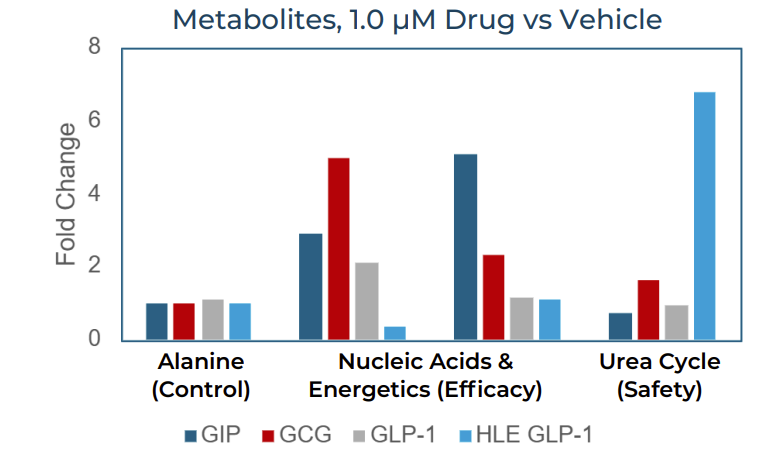
Pathway-level analysis of the 1.0 μM treatment data provided mechanistic resolution. Using alanine as a housekeeping metabolic control, Pyxis quantified fold-changes across three functionally informative pathway groups: nucleic acids and energetics (an indicator of target engagement and anabolic activity), and the urea cycle (a marker of cellular stress and potential cytotoxicity).
Fold-change in representative metabolite pathway groups at 1 μM drug vs. vehicle. GIP, GCG, and GLP-1 induced robust increases in nucleic acid and energetics metabolites (efficacy signal), while HLE GLP-1 failed to replicate this response and instead showed elevated urea cycle metabolites (safety signal).
The GIP, GCG, and GLP-1 control peptides all induced robust upregulation of nucleic acid and energetics pathway metabolites, consistent with productive receptor engagement and stimulated anabolic metabolism. In contrast, the HLE mimetic failed to elicit these expected efficacy signals. Simultaneously, the HLE candidate drove elevated urea cycle metabolite concentrations relative to all controls, indicating activation of stress-associated metabolic pathways potentially linked to cytotoxicity.
Latent-Space Metabolic Fingerprinting Provides Complementary Candidate Ranking
Beyond pathway-level quantitation, Pyxis encodes cellular metabolic state as a high-dimensional latent-space fingerprint derived from the full spectral profile. Dimensionality reduction via UMAP projects these fingerprints into interpretable two-dimensional space, enabling direct visual comparison of compound-induced metabolic states across all candidates simultaneously.
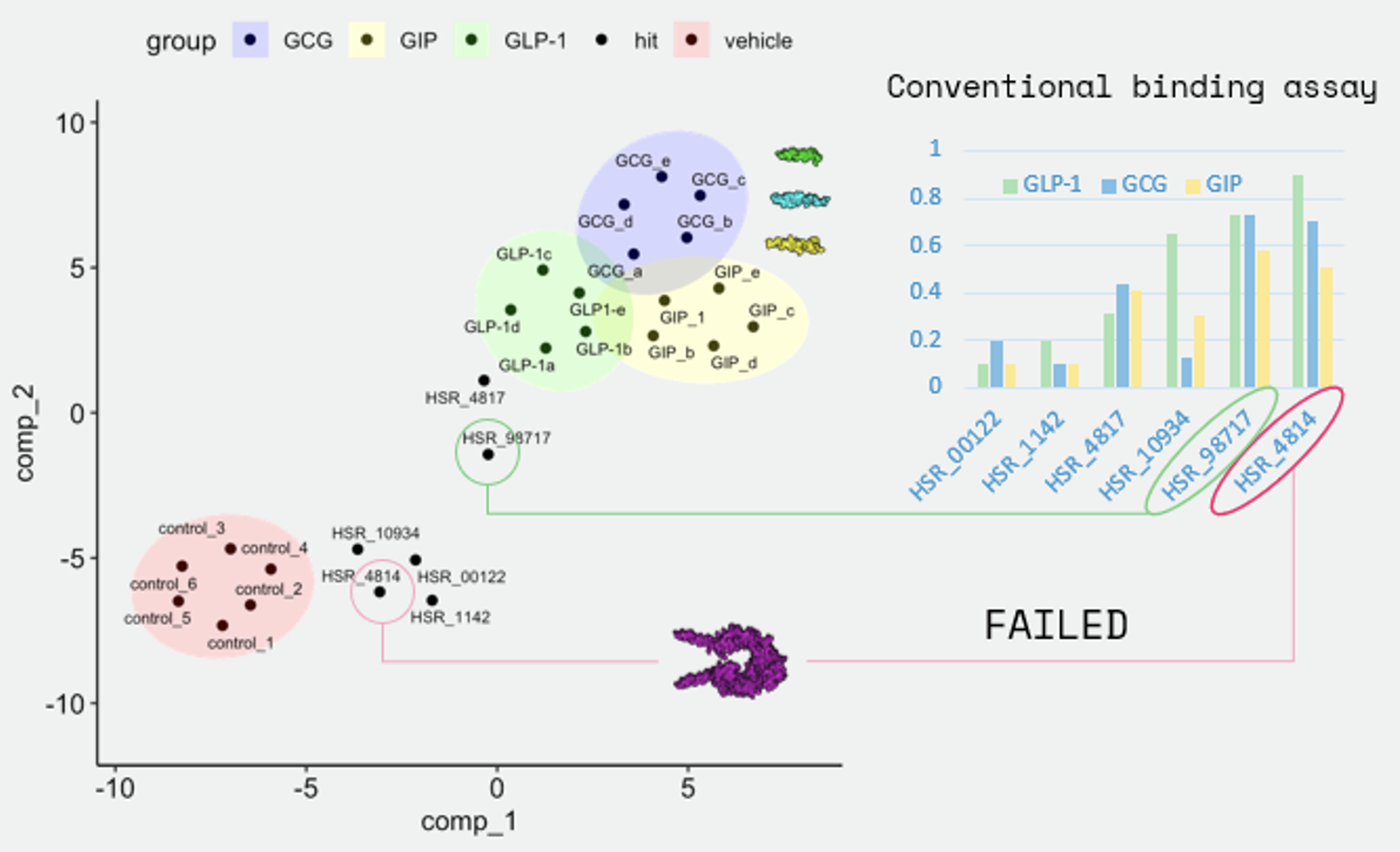
UMAP projection of Pyxis latent-space metabolic fingerprints for CFPAC-1 cells treated with each incretin mimetic. GLP-1, GCG, and GIP candidates form distinct but proximal clusters consistent with incretin receptor activation. The HLE candidate (labeled FAILED) projects to an isolated region of metabolic space, irrespective of its conventional receptor binding scores (inset bar chart), demonstrating that metabolic fingerprinting captures biological activity orthogonal to binding affinity.
In the UMAP projection, GLP-1, GCG, and GIP candidates form discrete but proximal clusters in a region of metabolic space consistent with productive incretin receptor activation. The HLE candidate, by contrast, projects to an isolated region of the manifold, clearly separated from all functional controls. Critically, this metabolic failure was not predicted by conventional receptor binding assays (shown in inset), which assigned the HLE candidate moderate-to-high binding scores for GLP-1, GCG, and GIP receptors. Pyxis latent-space fingerprinting thus revealed functional information inaccessible to affinity-based screening alone.
Discontinuation of the HLE candidate as a lead was recommended based on Pyxis metabolic data. This recommendation was subsequently corroborated by orthogonal functional assays conducted at a contract research organization (CRO), validating the predictive power of metabolomics-based triage.
AI-Powered In Vitro Metabolic Screening with Pyxis
Cellular metabolism provides a rich, multi-dimensional readout of drug action. Shifts in glycolysis, the TCA cycle, nucleotide biosynthesis, and amino acid metabolism collectively reflect whether a therapeutic candidate is engaging its intended target, activating off-target pathways, or causing cellular stress. Capturing these signals simultaneously — with quantitative accuracy and without prior knowledge of which metabolites to measure — requires a fundamentally different analytical approach.
Pyxis, built on a proprietary Large Spectral Model (LSM), converts raw LC-MS data into confident metabolite identities and absolute concentrations within minutes of data upload. Its standardized acquisition method and universal calibrator eliminate assay-specific method development, making it ideally suited to high-throughput screening workflows. Broad, unbiased metabolic coverage enables simultaneous assessment of receptor engagement (efficacy) and cell health (safety) from a single LC-MS run.
See what Pyxis finds in your data
Upload a dataset you've already analyzed. You'll see similar results, faster, and you may even learn something new.
Sign up for Pyxis