Try it yourself — no credit card required
Get annotations in minutes.
AI-powered metabolite identification. No scripting, no parameter tuning. Just drag, drop, and discover.
Try it on our demo datasets in under 10 minutes
Used by researchers at leading pharma, biotech, and academic institutions
1.8M reference spectra
99K unique analytes
Results in <10 minutes
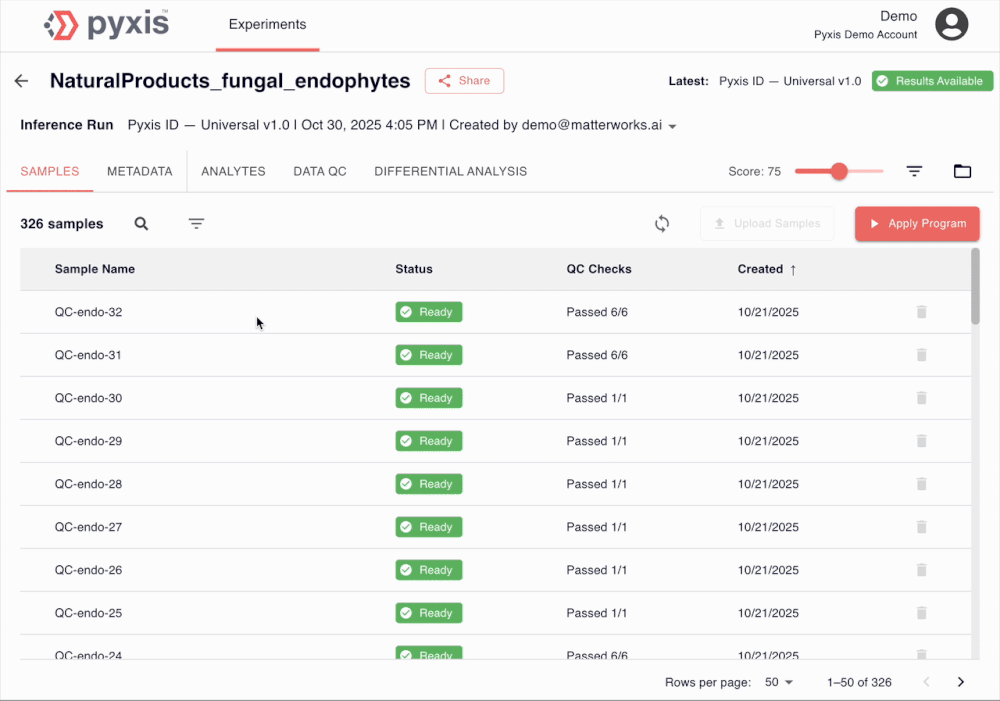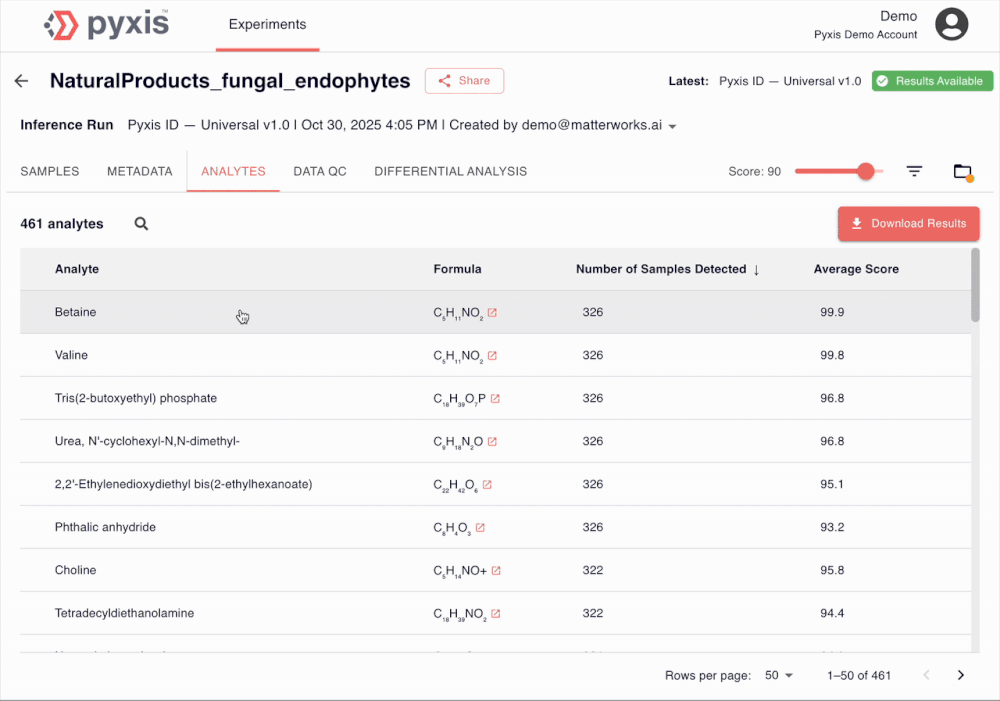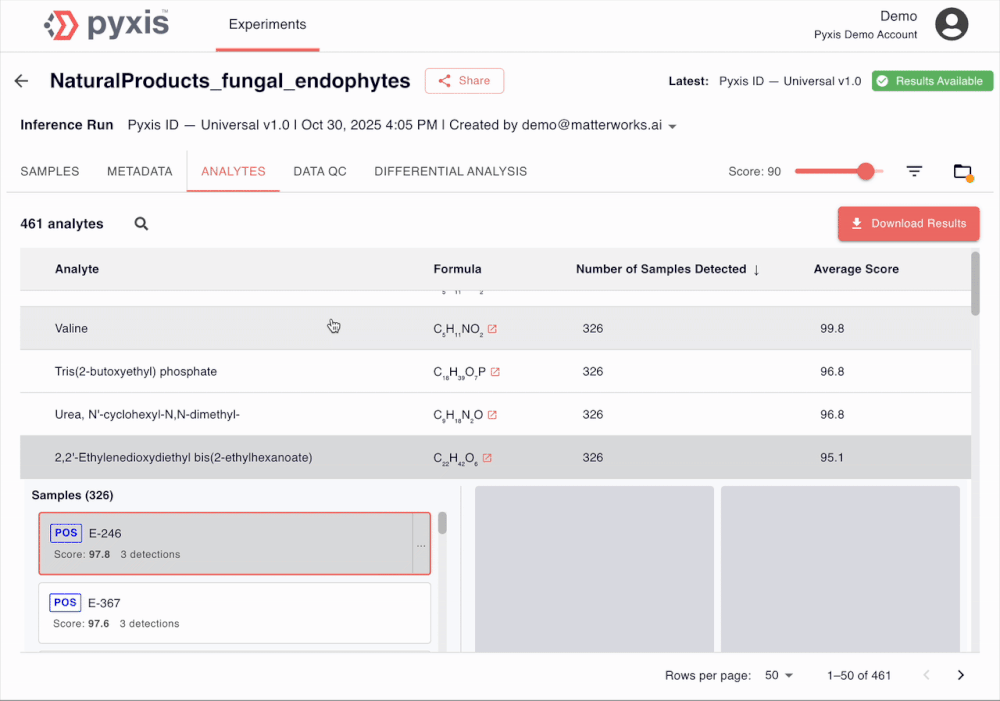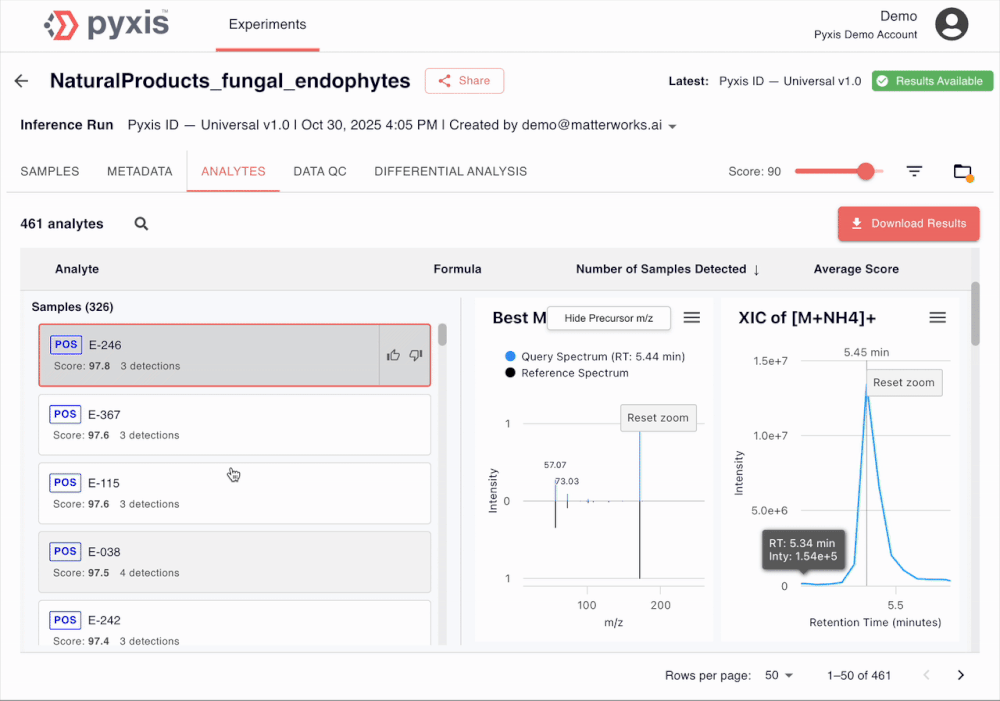
See Pyxis in action
A 5-minute walkthrough of a human disease plasma dataset with our Head of Biology. Sign up to see the demo dataset yourself.
How it works
From raw data to publication-ready annotations in three steps.
1
Upload your spectra
Drag and drop your MS2 files. We support .raw, .mzML, .mzXML, .wiff, and more. No preprocessing required.
2
AI identifies compounds
Our model searches 1.8M reference spectra to identify metabolites, lipids, and other biomolecules automatically.
5-10 minutes
3
Export your results
Review spectral similarity, examine spectra matches, and export annotations for your analysis pipeline.
Instant